The Biochemistry and Biophysics department has active research in molecular biophysics and structural biology on a wide range of topics including intrinsically disordered proteins (IDPs), the dynamics of molecular motors, membrane protein biophysics, RNA structure and function, and many facets of protein signaling. Our faculty use many techniques including nuclear magnetic resonance (NMR), single-molecular light microscopy, molecular simulation, and isothermal titration calorimetry.
Breadcrumb
- BB
- Back to Research
- Molecular Biophysics and Structural Biology
Molecular Biophysics and Structural Biology
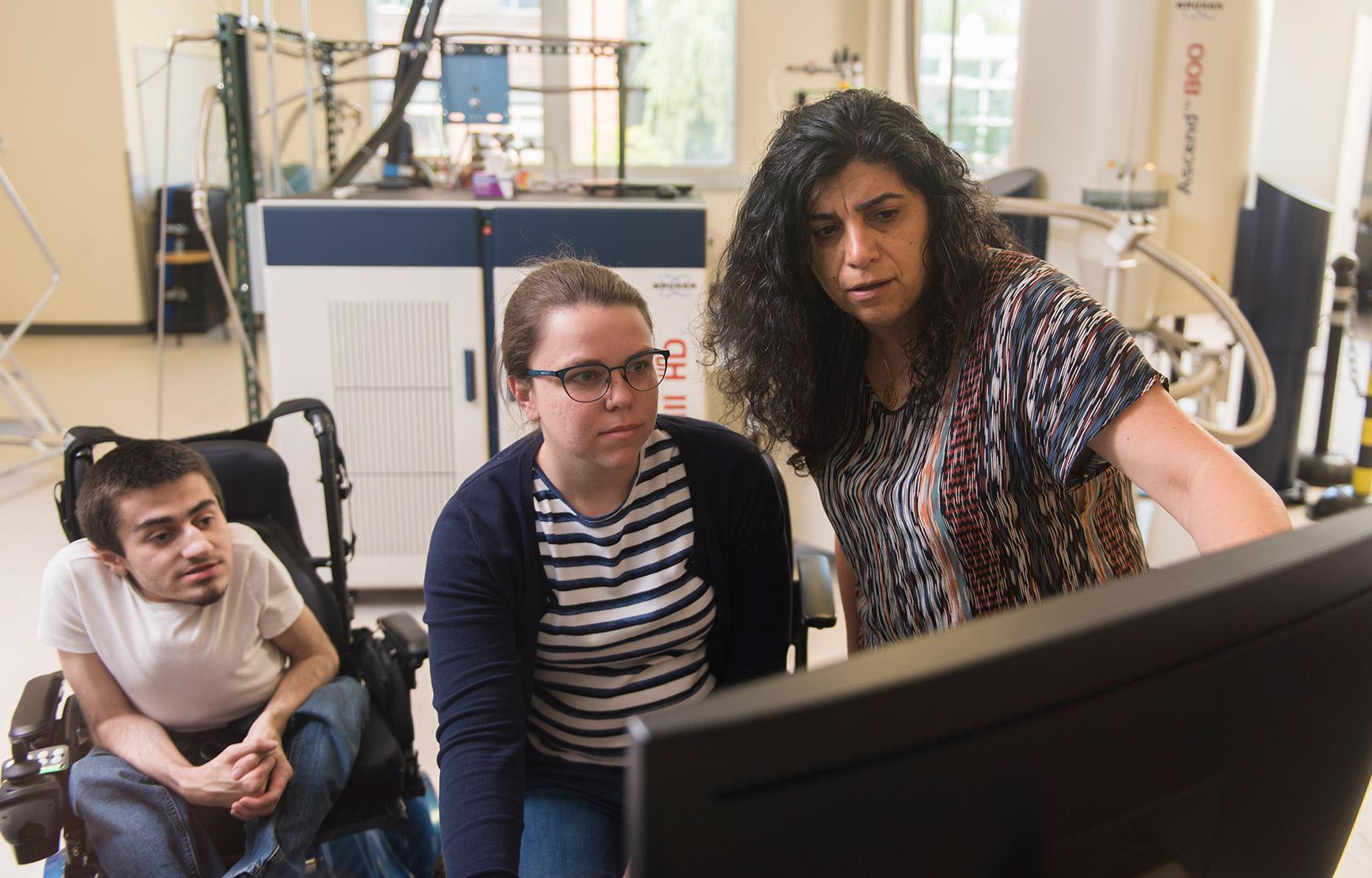

Molecular biophysics and structural biology researchers
Research faculty are accepting new graduate students unless designated with (*).

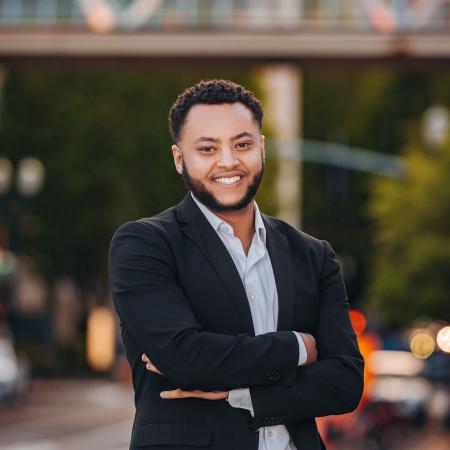
Elisar Barbar

Biochemistry and Biophysics Professor, Faculty Director of OSU's Biomolecular NMR Facility
Elisar Barbar
The Barbar lab focuses on elucidating molecular processes that govern protein networks involving intrinsically disordered proteins (IDPs). Our approach includes characterization of protein assemblies using combined analysis from multiple biophysical techniques. We obtain structural information by NMR, crystallography, and negative stain electron microscopy, binding thermodynamics by isothermal titration calorimetry (ITC), hydrodynamics by size exclusion chromatography multiangle light scattering (SEC-MALS) and analytical ultracentrifugation (AUC), and dynamics by NMR. Current projects focus on the intermediate chain of cytoplasmic dynein, viral proteins (Rabies, COVID, HPIV, Ebola), the transcription factor ASCIZ, a tumor suppressor 53BP1, and other partners of the hub protein LC8.
View her profile
Read more about the NMR Facility
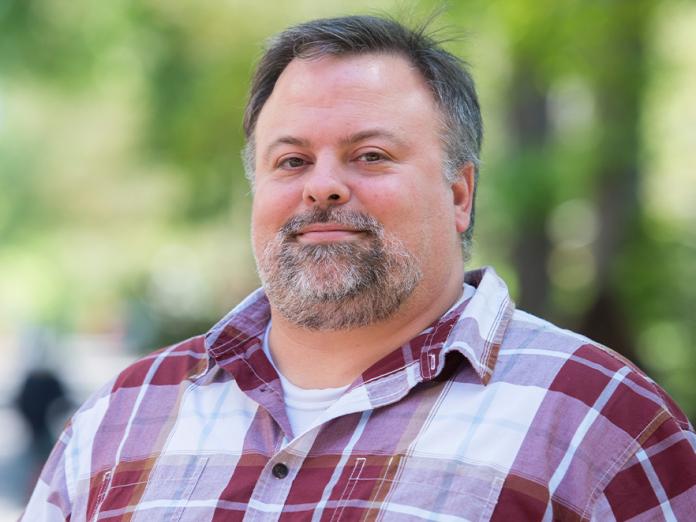
Afua Nyarko
Afua Nyarko
The Nyarko lab investigates mechanisms of multivalent protein assemblies in signaling pathways, with the goal of identifying novel strategies to selectively modulate downstream responses. Our primary focus is the Hippo signaling pathway, a network of protein-protein associations critical to the homeostatic balance of cell growth versus cell death. We combine a variety of mutually reinforcing molecular biophysics methodologies including solution NMR to advance understanding of the fundamental mechanisms of protein associations in the Hippo signaling pathway, elucidate the regulatory importance of multivalent intrinsically disordered proteins in the signaling network, and decipher the strong link between Hippo signaling dysregulation and cancer.
View her profile

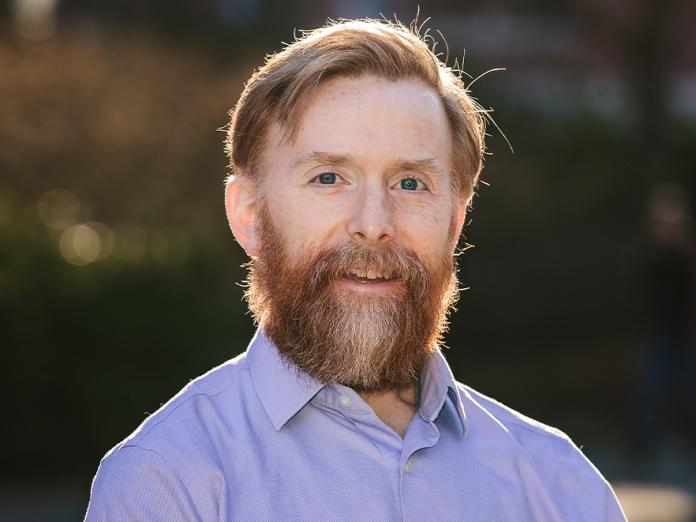
Juan M. Vanegas

Juan M. Vanegas
The Vanegas laboratory combines techniques from molecular simulation, continuum mechanics, and quantum chemistry to understand how molecular structure modulates the activation of mechanosensitive proteins and determines the mechanical response of lipid membranes. His group is highly interdisciplinary working at the interface between biology, physics, chemistry, and engineering. The central focus of our research is to provide mechanistic insights into essential biological processes such as membrane fission and fusion, organelle and cellular shaping, touch and pain sensing, cardiovascular control and development, and osmotic regulation among others. He works closely with other laboratories to integrate our modeling efforts with experimental results.
View his profile

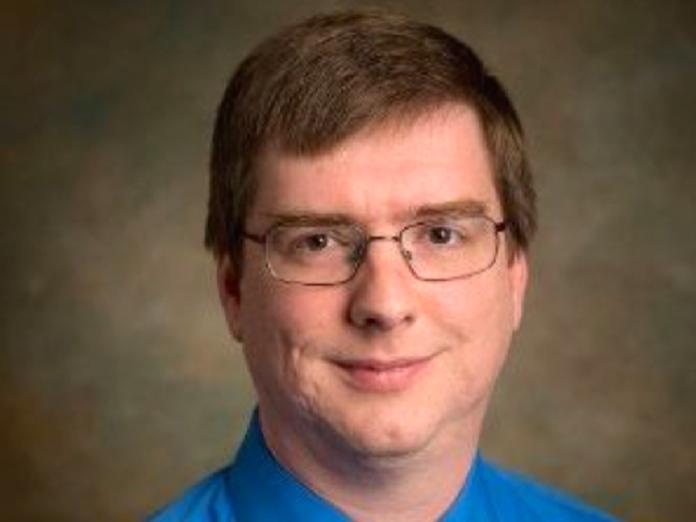
David Hendrix
The Hendrix lab has organized the largest meta-database of RNA secondary structures. This database has enabled the analysis of secondary structure, including uncovering patterns associated with specific structures, balance of thermodynamic energy, and in the development of new structural alignment methods. We have also developed several approaches including machine learning and deep learning to microRNA discovery and analysis of microRNA biogenesis.
View his profile

Colin Johnson
The Johnson lab studies the role of membrane proteins in Ca2+ signaling and disease. Through development of new organismal models for in vivo study of pathological mutations, and reconstitution of the signaling steps in an in vitro setting, we have identified key proteins that link rises in intracellular Ca2+ with physiological processes and dysfunction including hearing, deafness, cell membrane repair, and muscular dystrophy. Our long-term goals are both to improve our basic understanding of the signaling associated with these processes, and the development of potential therapeutics.
View his profile

Rick Cooley

Rick Cooley
The Cooley lab seeks to understand how phosphorylation regulates protein structure and function to coordinate cellular signaling events, with the ultimate goal of discovering novel strategies by which specific phosphorylated forms of proteins can be targeted for therapeutic intervention. Using a combination of Genetic Code Expansion and a variety of biophysical techniques including x-ray crystallography, small angle x-ray scattering and NMR, we focus on i) uncovering mechanisms by which 14-3-3 regulates phospho-protein function and ii) how phosphorylation alters protein-protein interactions in apoptotic pathways.
View his profile

Sarah Clark

Sarah Clark
What is the molecular basis of touch sensation? Organisms sense and interact with their environment through the perception of mechanical stimuli such as touch, vibration, temperature, and sound. Specialized sensory neurons in the skin and inner ear detect mechanical stimuli and convert it into an electrical signal through the opening of ion channels in the cell membrane. The Clark lab studies the architecture and mechanism of mechanically-activated ion channels to understand the molecular basis of touch sensation. We use C. elegans as a model system and apply a variety of biophysical and biochemical methods to our work, including cryo-electron microscopy (cryo-EM), fluorescence microscopy, TIRF microscopy, and patch-clamp electrophysiology.
How are lipids trafficked to the cell membrane? Lipids are synthesized in the endoplasmic reticulum (ER) and moved to different organelles via non-vesicular and vesicular trafficking. Non-vesicular lipid trafficking is carried out by lipid transport proteins, large macromolecular complexes that form a bridge between membranes at organelle-membrane contact sites. Lipid transport proteins are proposed to act as ‘lipid firehoses’, rapidly shuttling single lipids between donor and acceptor membranes. We use cryo-EM, as well as other biophysical and biochemical techniques, to study the architecture of lipid transport protein complexes with the goal of understanding the mechanism of lipid transport.

Dan Liefwalker

Assistant Professor
Dan Liefwalker
MYC is an intrinsically disordered protein lacking enzymatic regions as therapeutic targets. Sequential posttranslational modifications signal for MYC activity or degradation, which is tightly controlled in normal cells. The Liefwalker lab is investigating the role of MYC-dependent posttranslational modification in regulating the activity of histone demethylases that also act as scaffold proteins bound to histone complexes and how these interactions influence chromatin architecture.
View his profile

Patrick Reardon
Dr. Reardon directs the OSU NMR Facility and conducts research focused on using NMR to study complex biological problems. We are developing In-Cell NMR tools to enable atomic resolution structural and dynamic analysis of proteins in living cells. We also develop NMR methods to measure stable isotope labeling in metabolites and utilize this information to map metabolic fluxes in biological systems. Dr. Reardon operates the analytical ultracentrifuge and collaborates with a number of groups to characterize macromolecules and their complexes.
View his profile

Weihong Qiu
The Qiu lab seeks to understand the interplay between kinesin-5 and kinesin-14 motor proteins in bipolar spindle assembly using an interdisciplinary approach that integrates molecular biology, protein biochemistry, cell biology, structural biology, physics-based theoretical modeling, and single-molecule light microscopy.
View his profile

Andy Karplus*
Dr. Karplus no longer leads a research group but still is involved in collaborative work with others who study proteins. Proteins play central roles in all aspects of biochemistry and detailed structural information is a key ingredient to understanding the mechanisms involved in protein function. One main expertise Karplus brings is in solving detailed protein structures by X-ray crystallography, and in interpreting those structures in light of functional and evolutionary information to figure out how they work.
View his profile

Victor Hsu*
The Hsu laboratory is interested in studying the structural aspects of biomolecular recognition and interactions, especially in protein-nucleic acid complexes. These interactions account for many of the major cell functions such as the induction or repression of gene expression and the packaging of nucleic acids into other superstructures. The primary technique that he uses is nuclear magnetic resonance (NMR) spectroscopy, which is uniquely suited for studying biomolecular structures at atomic resolution. The lab studies both sequence-specific and nonspecific DNA-binding proteins and has been active in developing isotope-edited NMR strategies to obtain more accurate distance constraints for use in structure calculations and to investigate the intrinsic flexibility of protein and DNA backbones.
View his profile

Myriam Cotten

Associate Professor, OSU-Cascades Campus
Myriam Cotten
The Cotten lab investigates proteins and peptides that interact with lipids as part of host defense and molecular delivery. Specific molecular systems include host defense peptides and lipids, cell penetrating peptides, and lipid-transfer proteins. We establish structure-dynamics-activity relationships and develop new mechanistic understanding by using complementary biophysical and biochemical techniques such as nuclear magnetic resonance (NMR), circular dichroism, calorimetry, surface plasmon resonance, and activity assays on live cells and mimics of their plasma membranes. Through biomolecular engineering approaches, we design enhanced analogs. Current projects include establishing synergistic antimicrobial, antiviral, and anticancer effects between peptides and other agents such as glycolipids; characterizing the mechanisms of membrane crossing by direct delivery peptides and of lipid transport by lipid-transfer proteins; and leveraging high-field NMR to gain new insights into the interplay between the physicochemical properties of biological membranes and their functional roles.
View her profile

Liang Huang

Liang Huang
I was known as a computational linguist (parsing, translation, algorithms and theory), but in recent years I became more interested in adapting my NLP algorithms to computational biology, thanks to the shared mathematical foundations between the two seemingly distant fields. In particular, I have been working on efficient (mostly linear-time) algorithms for RNA folding, mRNA design, homologous folding, and RNA design. More interestingly, when COVID-19 hit, this line of work became much more relevant because SARS-CoV-2 is the longest RNA virus known today (~30,000 nucleotides) which requires linear runtime, and because mRNA vaccine is the best way to prevent it but mRNA design is computationally challenging. Therefore, my work has made impact on the fight against COVID-19, and has resulted in high-profile papers such as LinearTurboFold (PNAS 2021) and LinearDesign (Nature 2023).
I am most interested in the theoretical and algorithmic aspects of biology and language, and many of my NLP/bio papers draw unexpected connections from theoretical computer science, e.g., my synchronous binarization algorithm (binarizing a synchronous context-free grammar in linear-time) was inspired by Graham Scan for Convex Hull, my LinearDesign algorithm (optimal mRNA design) uses the intersection between context-free and regular languages, and my k-best parsing algorithms are often featured in my Algorithms courses.
On the other hand, my recent work in NLP pioneered the field of simultaneous translation, turning it from obscurity to spotlight. See my ACL 2019 Keynote and CVPR 2021 Invited Talk.
Listing of my papers on Google Scholar, Semantic Scholar, and a subset (top conferences) in csrankings.org.
View Their Profile
Home to the highest-field NMR in the state of Oregon
The OSU Nuclear Magnetic Resonance (NMR) facility is a campus-wide core facility dedicated to providing state-of-the-art NMR spectroscopy resources to the research and education community at OSU and throughout the Pacific Northwest. In an effort led by Elisar Barbar, more than 2.4 million in funding was secured for the 800 MHz instrument and supported the hire of facility director Patrick Reardon. The facility supports scientific inquiry in a diverse array of research areas such as structural biology, organic chemistry, natural products analysis and environmental studies.
Latest Stories
Across the Department

GCE4ALL leads global advancements in genetic code expansion for advanced therapies








